Beyond K-Means#
K-Means is a very useful starting point, but it is not the only clustering method. Two important alternatives are:
Hierarchical Clustering
DBSCAN
Hierarchical Clustering#
Hierarchical clustering builds a hierarchy of clusters instead of assigning all points at once.
Key Idea#
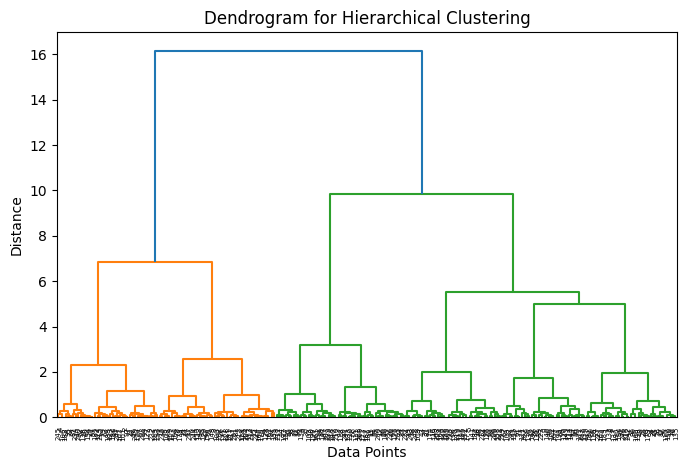
Instead of fixing the number of clusters beforehand, hierarchical clustering creates a tree-like structure of clusters. This structure is called a dendrogram
A dendrogram shows:
How clusters are merged or split
The distance at which this happens
By “cutting” the dendrogram at a certain height, we choose the number of clusters.
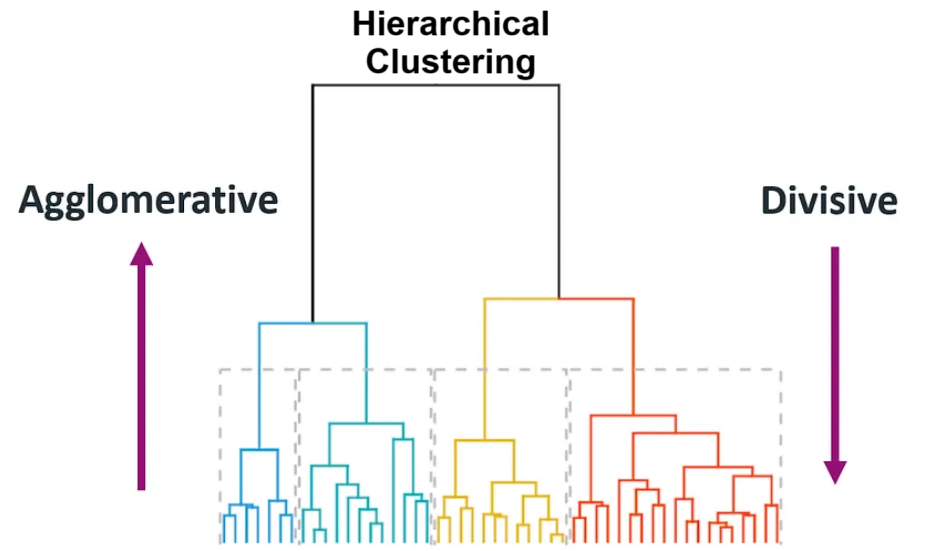
Figure: Dendrogram illustrating hierarchical clustering. The vertical axis shows the distance at which clusters are merged, and cutting the tree at different heights results in different numbers of clusters. Source: Medium
There are two common approaches:
(1) Agglomerative: (Start small → merge upward) Start with each point as its own cluster, then repeatedly merge the closest clusters.
(2) Divisive: (Start big → split downward) Start with one large cluster, then repeatedly split it
Agglomerative merges clusters step-by-step, while divisive splits clusters step-by-step, forming a hierarchy of clusters.
(1) Agglomerative (Bottom-Up): How it works:#
Start with individual points → merge clusters
Compute all distances → pick the smallest → merge → repeat
Algorithm Steps:#
Initialize:
Treat each data point as its own clusterCompute Distance Matrix:
Calculate pairwise distances between all clustersDistance between clusters depends on the linkage method:
Single Linkage: minimum distance between any two points
Complete Linkage: maximum distance between any two points
Average Linkage: average distance between all pairs
Ward’s Method: minimizes variance within clusters
The linkage method controls how clusters are merged, and can significantly affect the final clustering structure.

Figure: Comparison of linkage methods in hierarchical clustering: single, complete, average, and Ward’s method. Single linkage may create long chains, while complete linkage produces tighter and more compact clusters. Source: Analytics Vidhya
This step gives us a table of distances between every pair of clusters
Example: Distance Matrix
A |
B |
C |
|
|---|---|---|---|
A |
— |
2 |
5 |
B |
2 |
— |
3 |
C |
5 |
3 |
— |
Find Closest Clusters:
From the distance matrix, identify the two clusters with the smallest distanceExample: smallest distance = 2 (A–B)
So clusters A and B will be merged next
Merge Clusters:
Combine the selected clusters into a single clusterUpdate Distances:
Recompute distances between the new cluster and all others
(using the same linkage method)Repeat Steps 3–5
Continue until:Only one cluster remains OR
Desired number of clusters is reached
(2) Divisive (Top-Down): How it works:#
Start with one cluster → split into smaller clusters
Algorithm Steps:#
Initialize:
Put all data points into one clusterSelect Cluster to Split:
Choose the cluster that is most heterogeneous
(e.g., highest variance or largest size)Split Cluster:
Divide it into two smaller or sub clusters (often using a method like K-Means).cThe goal is to maximize separation between the two new clustersEvaluate Split:
Check if further splitting improves cluster quality (e.g., better separation or reduced variance).Repeat Steps 2–4
Continue until:Each point is its own cluster OR
Desired number of clusters is reached
Divisive clustering is less commonly used because splitting is computationally more complex than merging.
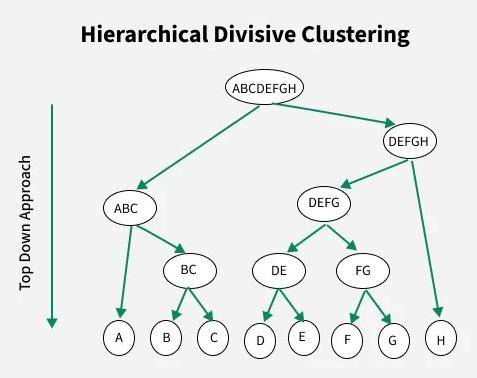
Figure: Divisive clustering process (top-down). The algorithm starts with one large cluster and recursively splits it into smaller clusters until the desired structure is obtained. Source: GeeksforGeeks
Why Use Hierarchical Clustering?#
No need to specify number of clusters initially
Provides insight into data structure
Useful for small to medium datasets
Limitation#
Computationally expensive for large datasets
Decisions (merge/split) cannot be undone
# Python Example
import matplotlib.pyplot as plt
from sklearn.datasets import make_moons
from sklearn.cluster import AgglomerativeClustering
from scipy.cluster.hierarchy import linkage, dendrogram
# Sample dataset
X, _ = make_moons(n_samples=250, noise=0.08, random_state=42)
# Apply Hierarchical Clustering
hc = AgglomerativeClustering(n_clusters=3, linkage='ward')
hc_labels = hc.fit_predict(X)
# -----------------------------
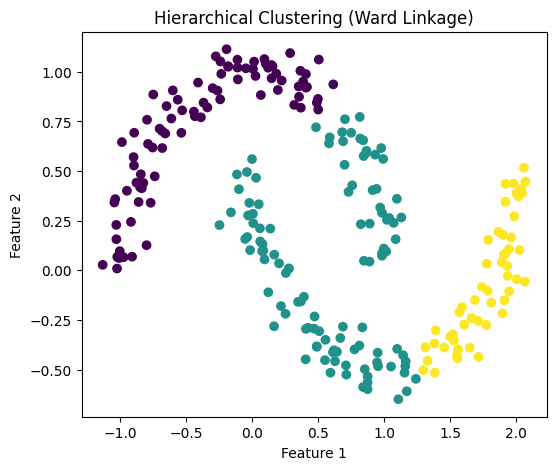
# Plot 1: Hierarchical Clusters
# -----------------------------
plt.figure(figsize=(6,5))
plt.scatter(X[:, 0], X[:, 1], c=hc_labels)
plt.xlabel("Feature 1")
plt.ylabel("Feature 2")
plt.title("Hierarchical Clustering (Ward Linkage)")
plt.show()
# -----------------------------
# Plot 2: Dendrogram
# -----------------------------
Z = linkage(X, method='ward')
plt.figure(figsize=(8,5))
dendrogram(Z)
plt.title("Dendrogram for Hierarchical Clustering")
plt.xlabel("Data Points")
plt.ylabel("Distance")
plt.show()
DBSCAN#
DBSCAN stands for: Density-Based Spatial Clustering of Applications with Noise. Unlike K-Means, DBSCAN does not try to create a fixed number of clusters. Instead, it groups together points that are packed closely together and labels isolated points as noise.
Main Idea#
Dense regions form clusters
Sparse regions may be treated as outliers
This makes DBSCAN especially useful when:
Clusters have irregular shapes
The dataset contains noise or outliers
Number of clusters is unknown
Key Parameters:#
DBSCAN uses two parameters: ε (neighborhood radius) and minPts (minimum points) to define dense regions and form clusters.
ε (epsilon): radius defining neighborhood
minPts: minimum number of points required to form a dense region
Types of Points#
Core Point: has at least
minPtsneighbors within εBorder Point: close to a core point but not dense enough itself
Noise Point: not close to any cluster
How DBSCAN Works#
Pick an unvisited point
Find all points within distance ε (epsilon)
This forms the point’s neighborhood
Check if the neighborhood has at least minPts points
If YES → it is a core point, start a new cluster
If NO → mark it as noise (may later become a border point)
Expand the cluster:
Add all points within ε of the core point
For each new point:
If it also has ≥ minPts neighbors → expand further
This creates a chain of density-connected points
Repeat until all reachable points are added
Move to next unvisited point and repeat
DBSCAN finds clusters based on density, grouping nearby points together while identifying sparse points as noise.
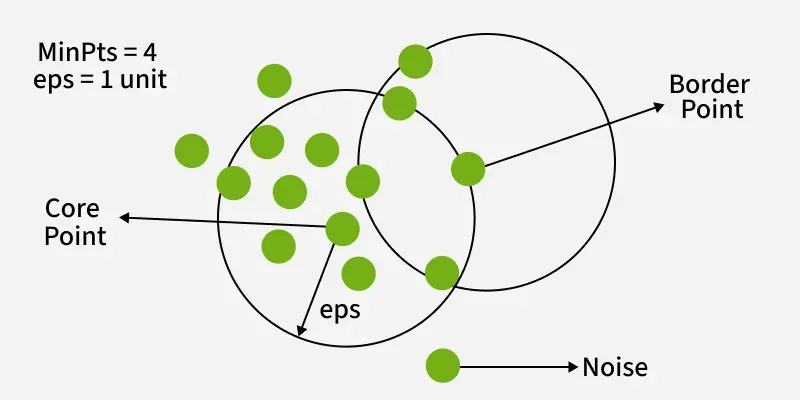
Figure: DBSCAN clustering showing core points (dense regions), border points (edges of clusters), and noise points (outliers). Source: GeeksforGeeks
In this figure:
Core points form the dense center of clusters
Border points lie on the edges of clusters
Noise points are isolated and do not belong to any cluster
DBSCAN builds clusters by expanding from core points and including all density-connected points (points reachable through neighboring core points).
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.animation import FuncAnimation
from IPython.display import HTML
from sklearn.datasets import make_moons
from sklearn.cluster import DBSCAN
from sklearn.neighbors import NearestNeighbors
# -----------------------
# Generate dataset
# -----------------------
X, _ = make_moons(n_samples=200, noise=0.08, random_state=42)
# Parameters
eps = 0.25
minPts = 5
# Pick a starting point
start_idx = 100
start_point = X[start_idx]
# Find neighbors within epsilon
nbrs = NearestNeighbors(radius=eps)
nbrs.fit(X)
neighbors = nbrs.radius_neighbors([start_point], return_distance=False)[0]
# Final DBSCAN result
db = DBSCAN(eps=eps, min_samples=minPts)
labels = db.fit_predict(X)
# -----------------------
# Set up figure
# -----------------------
fig, ax = plt.subplots(figsize=(6, 5))
def draw_base():
ax.clear()
ax.set_xlim(X[:, 0].min() - 0.5, X[:, 0].max() + 0.5)
ax.set_ylim(X[:, 1].min() - 0.5, X[:, 1].max() + 0.5)
ax.set_aspect("equal")
def update(frame):
draw_base()
if frame == 0:
ax.scatter(X[:, 0], X[:, 1], alpha=0.7)
ax.set_title("Step 1: Original Data")
elif frame == 1:
ax.scatter(X[:, 0], X[:, 1], alpha=0.7)
ax.scatter(start_point[0], start_point[1], s=120, marker='x')
ax.set_title("Step 2: Select an Unvisited Point")
elif frame == 2:
ax.scatter(X[:, 0], X[:, 1], alpha=0.7)
ax.scatter(start_point[0], start_point[1], s=120, marker='x')
circle = plt.Circle((start_point[0], start_point[1]), eps, fill=False, linewidth=2)
ax.add_patch(circle)
ax.set_title("Step 3: Find Points Within ε")
elif frame == 3:
ax.scatter(X[:, 0], X[:, 1], alpha=0.25)
ax.scatter(X[neighbors, 0], X[neighbors, 1], alpha=0.95)
ax.scatter(start_point[0], start_point[1], s=120, marker='x')
circle = plt.Circle((start_point[0], start_point[1]), eps, fill=False, linewidth=2)
ax.add_patch(circle)
ax.set_title(f"Step 4: Neighborhood Size = {len(neighbors)}")
elif frame == 4:
ax.scatter(X[:, 0], X[:, 1], alpha=0.25)
ax.scatter(X[neighbors, 0], X[neighbors, 1], alpha=0.95)
ax.scatter(start_point[0], start_point[1], s=120, marker='x')
circle = plt.Circle((start_point[0], start_point[1]), eps, fill=False, linewidth=2)
ax.add_patch(circle)
if len(neighbors) >= minPts:
ax.set_title(f"Step 5: Core Point ({len(neighbors)} ≥ {minPts}) → Start Cluster")
else:
ax.set_title(f"Step 5: Not a Core Point ({len(neighbors)} < {minPts}) → Noise/Border")
elif frame == 5:
unique_labels = sorted(set(labels))
for label in unique_labels:
if label == -1:
ax.scatter(
X[labels == label, 0],
X[labels == label, 1],
color='black',
label='Noise'
)
else:
ax.scatter(
X[labels == label, 0],
X[labels == label, 1],
label=f'Cluster {label}'
)
ax.legend()
ax.set_title("Final DBSCAN Clustering Result")
anim = FuncAnimation(fig, update, frames=6, interval=1800, repeat=True)
plt.close(fig)
HTML(anim.to_jshtml())
Why use DBSCAN?#
Does not require specifying the number of clusters in advance
Can detect outliers
Works well for arbitrarily shaped clusters
##Limitation Sensitive to parameter choices such as eps and min_samples
Comparing the Methods#
K-Means
Best for compact, spherical clusters
Requires choosing K
Hierarchical Clustering
Builds a tree-like clustering structure
Useful for exploring relationships between clusters
DBSCAN
Density-based
Handles noise and irregular cluster shapes better