import networkx as nx
import matplotlib.pyplot as plt
import numpy as np
Representing Graphs: 3 Ways to Store a Graph#
Before running any algorithm, you must choose a data structure (because you need to store the graph). The right choice depends on the graph’s size, density, and what operations you need to perform fast.
Summary#
Property |
Adjacency Matrix |
Adjacency List |
Adjacency Dictionary |
|---|---|---|---|
Space |
O(V²) |
O(V + E) |
O(V + E) |
Edge lookup |
O(1) |
O(degree) |
O(1) average |
Iterate neighbors |
O(V) |
O(degree) |
O(degree) |
Best for |
Dense graphs, fast lookups |
Sparse graphs (most real-world) |
Fast lookups with flexible structure |
Worst for |
Large sparse graphs |
Edge weight matrix |
Slight overhead due to hashing |
(1) Adjacency Matrix#
An N × N matrix where M[i][j] = 1 (or the edge weight) if an edge exists from node i to node j, otherwise 0.
Edge existence check in O(1)
Wastes O(V²) memory — 1 million nodes = 10¹² entries
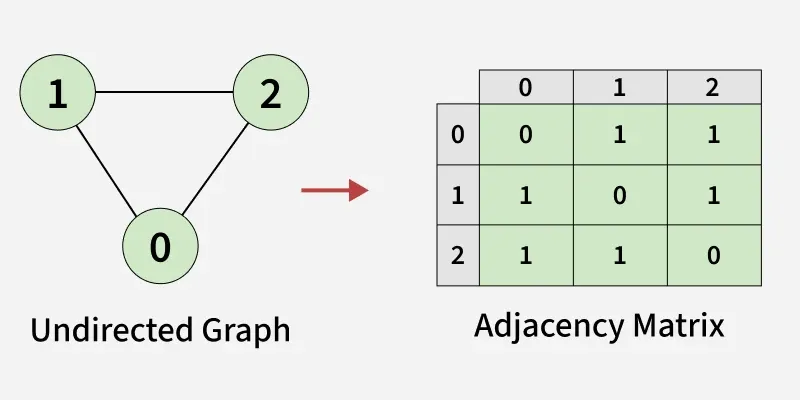
Adjacency Matrix. Source: GeeksforGeeks
# 4 nodes: You=0, Alice=1, Bob=2, Charlie=3
adj_matrix = [
[0, 1, 0, 1], # You connects to Alice and Charlie
[1, 0, 1, 1], # Alice connects to You, Bob, Charlie
[0, 1, 0, 0], # Bob connects to Alice
[1, 1, 0, 0], # Charlie connects to You and Alice
]
(2) Adjacency List#
Each node stores a list of its neighbors. Only edges that exist take up memory.
Space efficient: O(V + E) - far more efficient for sparse graphs (most real-world graphs have far fewer edges than the maximum possible).
Fast neighbor iteration — most graph algorithms use this
Checking if a specific edge exists requires scanning the neighbor list: O(degree)
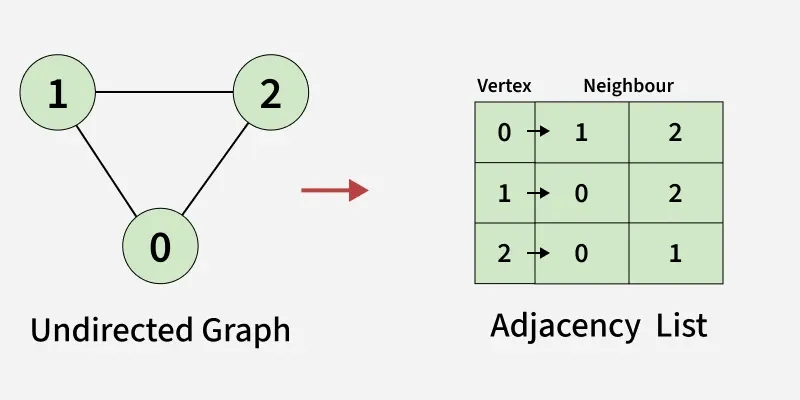
Adjacency List. Source: GeeksforGeeks
graph = {
'You': ['Alice', 'Charlie'],
'Alice': ['You', 'Bob', 'Charlie'],
'Bob': ['Alice', 'David'],
'Charlie': ['You', 'Alice', 'Eve'],
'David': ['Bob'],
'Eve': ['Bob', 'Charlie'],
}
Real-world graphs use libraries like NetworkX (Python), igraph, or graph databases like Neo4j for billion-edge scale.
(3) Adjacency Dictionary#
An adjacency dictionary is a dict-of-dicts. It stores each node and its neighbors as key–value pairs.
Outer key → node
Inner key → neighbor
Inner value → weight (or 1 if unweighted)
Some key advantages:
Fast edge lookup → average O(1)
Naturally supports weighted graphs
Flexible for storing extra information (distance, cost, frequency)
Intuition: Adjacency List = simple neighbors ; Adjacency Dictionary = neighbors + extra information
graph_weighted = {
'You': {'Alice': 2, 'Charlie': 1},
'Alice': {'You': 2, 'Bob': 3, 'Charlie': 2},
'Bob': {'Alice': 3, 'David': 4},
'Charlie': {'You': 1, 'Alice': 2, 'Eve': 5},
'David': {'Bob': 4},
'Eve': {'Bob': 2, 'Charlie': 5},
}
Compared to an adjacency list (dict of lists):
Operation |
Adj. List |
Adj. Dictionary |
|---|---|---|
Space |
O(V + E) |
O(V + E) |
Check if edge (u,v) exists |
O(degree(u)) |
O(1) |
Get weight of edge (u,v) |
O(degree(u)) |
O(1) |
Add edge |
O(1) append |
O(1) assign |
Remove edge |
O(degree) |
O(1) del |
Iterate neighbors of u |
O(degree) |
O(degree) |
This is the representation used internally by NetworkX (
G._adj) and by most production graph libraries.
Variants:
Unweighted — inner value is
1(or{}) — just tracks existenceWeighted — inner value is a number (distance, cost, probability)
Attributed — inner value is a dict holding multiple properties per edge
# ── Adjacency Matrix ─────────────────────────────────────────────────────────
node_list = ['You', 'Alice', 'Bob', 'Charlie', 'David', 'Eve']
n = len(node_list)
idx = {name: i for i, name in enumerate(node_list)}
adj_matrix = np.zeros((n, n), dtype=int)
for u, v in edges:
adj_matrix[idx[u]][idx[v]] = 1
adj_matrix[idx[v]][idx[u]] = 1 # undirected
df_matrix = pd.DataFrame(adj_matrix, index=node_list, columns=node_list)
print("=== Adjacency Matrix ===")
print(df_matrix)
# Visualise as heatmap
fig, ax = plt.subplots(figsize=(6, 5))
im = ax.imshow(adj_matrix, cmap='Blues', vmin=0, vmax=1)
ax.set_xticks(range(n)); ax.set_yticks(range(n))
ax.set_xticklabels(node_list, rotation=45, ha='right', fontsize=10)
ax.set_yticklabels(node_list, fontsize=10)
for i in range(n):
for j in range(n):
ax.text(j, i, adj_matrix[i][j], ha='center', va='center',
fontsize=12, fontweight='bold',
color='white' if adj_matrix[i][j] else '#cccccc')
ax.set_title('Adjacency Matrix', fontsize=13, fontweight='bold', pad=12)
plt.tight_layout()
plt.show()
sparsity = 1 - (adj_matrix.sum() / adj_matrix.size)
print(f"\nMatrix size : {n}×{n} = {n*n} cells")
print(f"Non-zero : {int(adj_matrix.sum())} (edges × 2 for undirected)")
print(f"Sparsity : {sparsity:.1%} ← most cells are wasted zeros")
=== Adjacency Matrix ===
You Alice Bob Charlie David Eve
You 0 1 0 1 0 0
Alice 1 0 1 1 0 0
Bob 0 1 0 0 1 1
Charlie 1 1 0 0 0 1
David 0 0 1 0 0 0
Eve 0 0 1 1 0 0
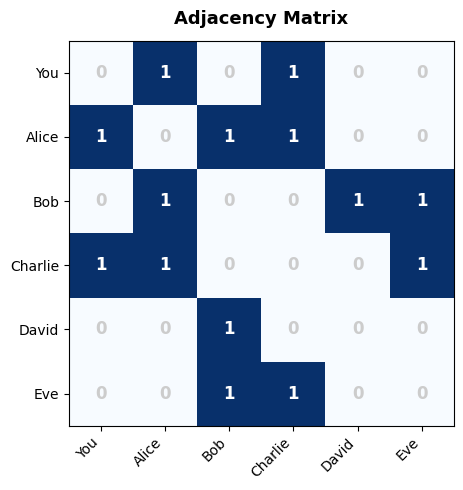
Matrix size : 6×6 = 36 cells
Non-zero : 14 (edges × 2 for undirected)
Sparsity : 61.1% ← most cells are wasted zeros
# ── Adjacency List ───────────────────────────────────────────────────────────
adj_list = {node: [] for node in node_list}
for u, v in edges:
adj_list[u].append(v)
adj_list[v].append(u) # undirected
print("=== Adjacency List ===")
for node, neighbors in adj_list.items():
print(f" {node:8s} → {neighbors}")
total_entries = sum(len(v) for v in adj_list.values())
print(f"\nTotal entries stored: {total_entries} (= 2×|E| = 2×{len(edges)})")
print(f"vs adjacency matrix : {n*n} entries")
print(f"Memory saving : {1 - total_entries/(n*n):.1%} fewer entries")
=== Adjacency List ===
You → ['Alice', 'Charlie']
Alice → ['You', 'Bob', 'Charlie']
Bob → ['Alice', 'David', 'Eve']
Charlie → ['You', 'Alice', 'Eve']
David → ['Bob']
Eve → ['Bob', 'Charlie']
Total entries stored: 14 (= 2×|E| = 2×7)
vs adjacency matrix : 36 entries
Memory saving : 61.1% fewer entries
# ── Adjacency Dictionary — three flavours ────────────────────────────────────
# 1. Unweighted (value = 1, just marks edge existence)
adj_dict_unweighted = {
'You': {'Alice': 1, 'Charlie': 1},
'Alice': {'You': 1, 'Bob': 1, 'Charlie': 1},
'Bob': {'Alice': 1, 'David': 1, 'Eve': 1},
'Charlie': {'You': 1, 'Alice': 1, 'Eve': 1},
'David': {'Bob': 1},
'Eve': {'Bob': 1, 'Charlie': 1},
}
# 2. Weighted (value = edge weight)
adj_dict_weighted = {
'You': {'Alice': 0.9, 'Charlie': 0.7},
'Alice': {'You': 0.9, 'Bob': 0.5, 'Charlie': 0.8},
'Bob': {'Alice': 0.5, 'David': 0.6, 'Eve': 0.4},
'Charlie': {'You': 0.7, 'Alice': 0.8, 'Eve': 0.3},
'David': {'Bob': 0.6},
'Eve': {'Bob': 0.4, 'Charlie': 0.3},
}
# 3. Attributed (value = dict with multiple properties per edge)
adj_dict_attributed = {
'You': {'Alice': {'weight': 0.9, 'since': 2023, 'context': 'orientation'},
'Charlie': {'weight': 0.7, 'since': 2023, 'context': 'class'}},
'Alice': {'You': {'weight': 0.9, 'since': 2023, 'context': 'orientation'},
'Bob': {'weight': 0.5, 'since': 2023, 'context': 'introduction'},
'Charlie': {'weight': 0.8, 'since': 2023, 'context': 'class'}},
# ... (truncated for brevity)
}
# ── O(1) operations ───────────────────────────────────────────────────────────
g = adj_dict_weighted
print("=== O(1) edge operations ===")
print(f"Edge (You, Alice) exists? {'Alice' in g['You']}") # O(1)
print(f"Edge (You, David) exists? {'David' in g['You']}") # O(1)
print(f"Weight of (Alice, Bob): {g['Alice']['Bob']}") # O(1)
print(f"Degree of Alice: {len(g['Alice'])}") # O(1)
print("\n=== Adding and removing edges ===")
g['David']['Eve'] = 0.2 # add a new edge O(1)
print(f"After adding David-Eve: {list(g['David'].keys())}")
del g['David']['Eve'] # remove it O(1)
print(f"After removing David-Eve: {list(g['David'].keys())}")
print("\n=== Iterating neighbors ===")
for neighbor, weight in g['Alice'].items(): # O(degree)
print(f" Alice → {neighbor:8s} weight={weight}")
=== O(1) edge operations ===
Edge (You, Alice) exists? True
Edge (You, David) exists? False
Weight of (Alice, Bob): 0.5
Degree of Alice: 3
=== Adding and removing edges ===
After adding David-Eve: ['Bob', 'Eve']
After removing David-Eve: ['Bob']
=== Iterating neighbors ===
Alice → You weight=0.9
Alice → Bob weight=0.5
Alice → Charlie weight=0.8